Tip: after the article is written, the directory can be generated automatically. Please refer to the help document on the right for how to generate it
1, Data sorting
1. sort
- If the direction is not indicated in sort, it will be sorted in ascending order by default
- In multidimensional tensors, sorting is not in the global case, but in the smallest dimension
# One dimensional case a = tf.random.shuffle(tf.range(10)) print(a) print(tf.sort(a)) print(tf.sort(a, direction='DESCENDING')) # Multidimensional situation b = tf.random.uniform([2, 4]) print(b) print(tf.sort(b)) print(tf.sort(b, direction='DESCENDING'))
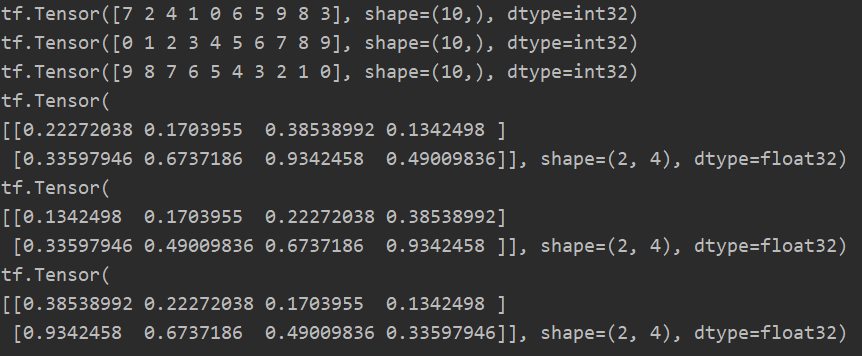
2. argsort
The returned value is not the sorted value, but the sorted index
a = tf.random.shuffle(tf.range(10)) print(a) print(tf.argsort(a)) print(tf.argsort(a, direction='DESCENDING')) # Multidimensional situation b = tf.random.uniform([2, 4]) print(b) print(tf.argsort(b))
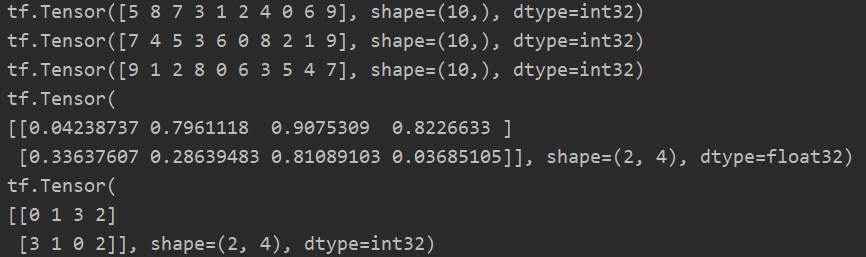
3. top_k
Returns the binary group of the first k largest elements, one is value and the other is index
c = tf.random.uniform([2, 5], maxval=20, dtype=tf.int32) print(c) res = tf.math.top_k(c, 3) print(res.indices) print(res.values)

2, Data segmentation
1. unstack
unstack is the inverse operation of stack, which separates a tensor according to a certain dimension
The code is as follows (example):
p = tf.ones([2, 4, 35, 8]) q, r = tf.unstack(p, axis=0) print(q.shape, r.shape)
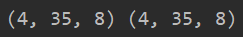
Here is a question: if axis is set to 3, that is to say, the dimension of 8 is separated. Can it be divided into two?
The answer is No. if you segment according to the dimension of 8, 8 new tensors will be generated, and the shape of each tensor is [2, 4, 35]
This is the problem of unstack, which cannot specify how many shares to divide.
p = tf.ones([2, 4, 35, 8]) res = tf.unstack(p, axis=3) print(len(res)) print(res[0].shape)
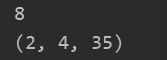
After unstack segmentation, the dimension of the data shape will be automatically reduced and the dimension of 1 will be removed
2. split
split is improved on the basis of unstack. You can specify any number of cutting copies and control the size of each copy.
The code is as follows (example):
x = tf.ones([2, 4, 35, 8]) # num_or_size_splits specifies how many pieces to cut into lips1 = tf.split(x, axis=3, num_or_size_splits=2) lips2 = tf.split(x, axis=3, num_or_size_splits=8) print(len(lips1)) print(lips1[0].shape) print(len(lips2)) print(lips2[0].shape) # You can also specify the size of each cut on the axis axis lips3 = tf.split(x, axis=3, num_or_size_splits=[2, 2, 4]) print(len(lips3)) print(lips3[0].shape) print(lips3[1].shape) print(lips3[2].shape)
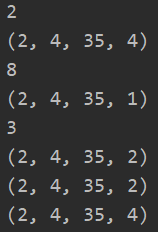
Even if the value of the dimension is 1, split will not automatically reduce the dimension
3, Data statistics
1. Norm
- One norm: take the absolute value of each element in the tensor and then sum it
- Two norm: the square of each element in the tensor, then the sum, and finally the root sign
- Infinite norm: take the absolute value of each element in the tensor, and then take the value with the largest absolute value
tf.norm function is used to find norm
- Set ord=1 to find L1 norm, and make ord=2 to find L2 norm. If ord is not given, L2 norm will be found by default
- Setting the axis value indicates that norm calculation is performed on a certain dimension
The code is as follows (example):
a = tf.ones([2, 2]) print(tf.norm(a)) print(tf.norm(a, ord=1)) print(tf.norm(a, ord=1, axis=1)) b = tf.ones([4, 28, 28, 3]) print(tf.norm(b))

2. Maximum value minimum value mean value
p = tf.random.normal([4, 10]) # Find the minimum value print(tf.reduce_min(p)) # When axis is not given, the minimum value of the whole tensor is obtained by default print(tf.reduce_min(p, axis=1)) # Find the minimum value on the axis with axis=1. Because there are four 10 columns, four data will be returned # Find the maximum value print(tf.reduce_max(p)) print(tf.reduce_max(p, axis=1)) # Find the maximum value on the axis with axis=1. Because there are four 10 columns, four data will be returned # Find the mean print(tf.reduce_mean(p)) print(tf.reduce_mean(p, axis=1)) # Calculate the mean value on the axis with axis=1. Because there are four 10 columns, four data will be returned
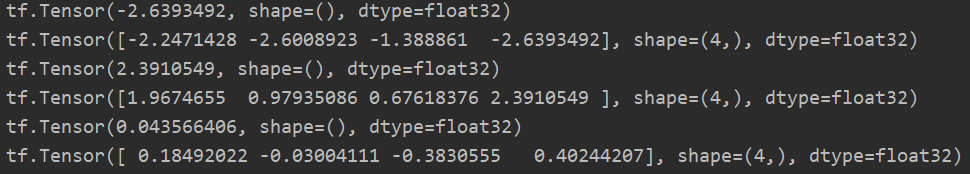
For the maximum and minimum values, there are also argmax and argmin functions to return the location information of the target value.
p = tf.random.normal([4, 10]) print(tf.argmin(p)) # When axis is not specified, the default axis is 0, so 10 data will be returned, and each data represents the index of the smallest element in each column print(tf.argmin(p, axis=1).shape) # axis=1, then there are 4 returned numbers print(tf.argmax(p)) print(tf.argmax(p, axis=1).shape)
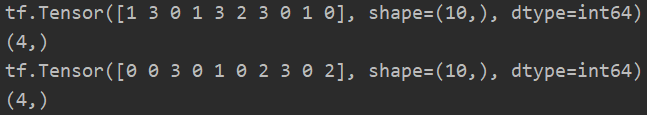
3. Comparison
tf.equal is used to compare the size relationship between tensors, provided that the shape s of the two tensors are the same. The comparison method is to compare the corresponding elements of each group in turn to judge whether they are equal. If they are equal, they are recorded as True and if they are unequal, they are recorded as False.
The return value is a truth table corresponding to the shape.
m = tf.random.normal([4, 10]) n = tf.random.normal([4, 10]) print(tf.equal(m, n))
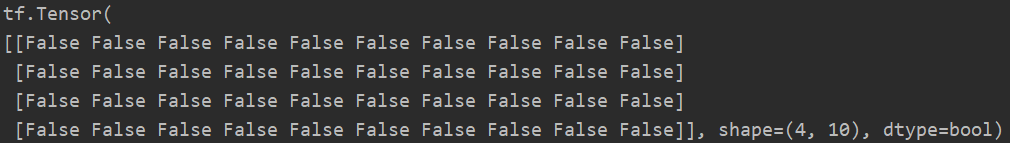
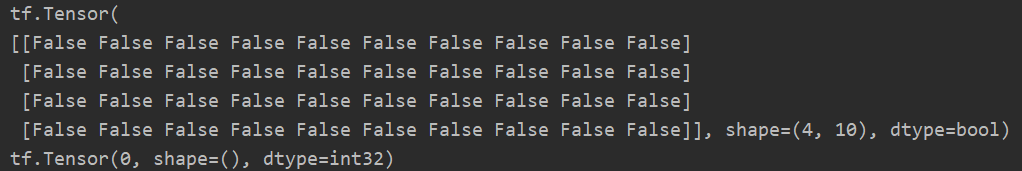
Using this method, we can judge how many predicted values are equal to the real values, that is, Accuracy can be calculated. Generally, if True=1 and False=0, the truth table can be transformed from bool type to int type, and then sum the whole table. The sum value represents how many values are equal.
m = tf.random.normal([4, 10]) n = tf.random.normal([4, 10]) print(tf.equal(m, n)) res = tf.equal(m, n) sum = tf.reduce_sum(tf.cast(res, dtype=tf.int32)) print(sum)
4. Weight removal
tf. The unique function returns an element that is not repeated in the tensor and records the position of the new element for the first time
x = tf.range(5) print(tf.unique(x)) y = tf.constant([4, 2, 3, 2, 3, 6]) print(tf.unique(y))

The output of unique is divided into two parts:
- The first part is the data after de duplication
- The second part is the mapping of the position of the data before operation and after de duplication