Summary of relevant codes of machine learning
Summary of machine learning related codes
XGBoost
There is one in sklearn, but there is another one with more powerful functions, as long as
pip3 install xgboost
Can be installed, but this installation process is really twists and turns.
Then, we need to know the general process used by xgboost. The following examples do not leave the process framework:
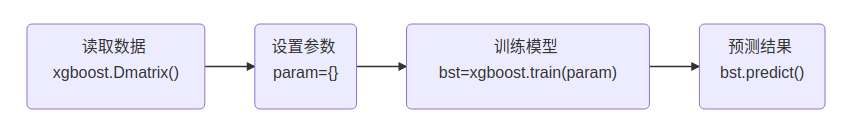
Example 1
In this example, an agaricus data is involved. This word means: Brazilian mushroom. But there are many kinds of Brazilian mushrooms, some toxic and some non-toxic. Can you predict whether a given mushroom is toxic or non-toxic?
import xgboost as xgb
import numpy as np
# 1. Basic use of xgBoost
# 2. Gradient and second derivative of custom loss function
train_data = 'xgboost_data/agaricus_train.txt'
test_data = 'xgboost_data/agaricus_test.txt'
# Define a loss function
def log_reg(y_hat, y):
p = 1.0 / (1.0 + np.exp(-y_hat))
g = p - y.get_label()
h = p * (1.0 - p)
return g, h
# Error rate. In this example, the estimated value < 0.5 means there is no poison
def error_rate(y_hat, y):
return 'error', float(sum(y.get_label() != (y_hat > 0.5))) / len(y_hat)
if __name__ == "__main__":
# Read data
data_train = xgb.DMatrix(train_data)
data_test = xgb.DMatrix(test_data)
# Set parameters
param = {'max_depth': 3, 'eta': 1, 'silent': 1, 'objective': 'binary:logistic'} # logitraw
watchlist = [(data_test, 'eval'), (data_train, 'train')]
n_round = 7
bst = xgb.train(param, data_train, num_boost_round=n_round, evals=watchlist, obj=log_reg, feval=error_rate)
# Calculation error rate
y_hat = bst.predict(data_test)
y = data_test.get_label()
print('y_hat',y_hat)
print('y', y)
error = sum(y != (y_hat > 0.5))
error_rate = float(error) / len(y_hat)
print('Total number of samples:\t', len(y_hat))
print('Number of errors:\t%4d' % error)
print('Error rate:\t%.5f%%' % (100 * error_rate))
explain:
Log defined at the beginning_ REG and error_ Finally, rate will be used in the following train method, corresponding to the obj parameter and the interval parameter, which means: use the user-defined loss function log_reg, to improve. Adopt user-defined error rate error_rate to predict the error rate.
About the train function:
def train(params, dtrain, num_boost_round=10, evals=(), obj=None, feval=None,
maximize=None, early_stopping_rounds=None, evals_result=None,
verbose_eval=True, xgb_model=None, callbacks=None)
"""
dtrain:Training data
num_boost_round:Number of iterations during data promotion
evals:Verify and pass in a tuple, which specifies what is the training set and what is the test set
"""
There is a params in the train, which involves the Booster parameter
-
max_depth: Specifies the depth of the decision tree
-
eta: learning rate, default 0.1
-
Silent: silent mode. If the value is 1, the model will not be output during operation
-
objective: given the loss function, the default is binary:logistic, or reg:linear
In xgboost, relevant data will be stored in DMatrix data structure, which is a two-dimensional matrix, but xgboost optimizes it.
Get keeps appearing in the above code_ Label method, so what is label?
There is a clear explanation in English:
The label is the name of some category. If you're building a machine learning system to distinguish fruits coming down a conveyor belt, labels for training samples might be "apple", " orange", "banana". The features are any kind of information you can extract about each sample. In our example, you might have one feature for colour, another for weight, another for length, and another for width. Maybe you would have some measure of concavity or linearity or ball-ness.
That is, when implemented in use, the label represents what you are at last, and the two feature features represent which attributes
Example 2
In this example, the iris dataset is used. In fact, there are many kinds of iris. For different iris (there are three types in this data set: setosa, versicolor and Virginia). Different species of iris have different attributes such as flower width and leaf length. Let's train with XGBoost to see if we can effectively predict the relevant data.
import numpy as np
import pandas as pd
import xgboost as xgb
from sklearn.model_selection import train_test_split # cross_validation
def iris_type(s):
it = {'Iris-setosa': 0, 'Iris-versicolor': 1, 'Iris-virginica': 2}
return it[s]
if __name__ == "__main__":
path = 'xgboost_data/iris.data' # Data file path
data = pd.read_csv(path, header=None)
x, y = data[range(4)], data[4]
y = pd.Categorical(y).codes
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=1, test_size=50)
data_train = xgb.DMatrix(x_train, label=y_train)
data_test = xgb.DMatrix(x_test, label=y_test)
watch_list = [(data_test, 'eval'), (data_train, 'train')]
#The depth of the decision tree is 2 and the learning rate is 0.3,
param = {'max_depth': 2, 'eta': 0.3, 'silent': 1, 'objective': 'multi:softmax', 'num_class': 3}
bst = xgb.train(param, data_train, num_boost_round=6, evals=watch_list)
y_hat = bst.predict(data_test)
result = y_test.reshape(1, -1) == y_hat
print('Correct rate:\t', float(np.sum(result)) / len(y_hat))
print('END.....\n')
explain:
- There is a PD in this code Categorica method. This method has the function of classification and sorting.
pandas.Categorical(val,category = None,ordered = None,dtype = None) """ val :[list-like] The values of categorical. categories:[index like] Unique categorisation of the categories. ordered :[boolean] If false, then the categorical is treated as unordered. dtype :[CategoricalDtype] an instance. Error- ValueError: If the categories do not validate. TypeError : If an explicit ordered = True but categorical can't be sorted. Return- Categorical varibale """
-
[reshape(1,-1) converted to 1 line
[reshape(2,-1) converted to two lines
[reshape(-1,1) converted to 1 column
[reshape(-1,2) converted to two columns