Computer vision life learn SLAM learning notes from scratch
The following topics are from computer vision life. Learn SLAM series from scratch
Point cloud fusion
subject
Title: point cloud fusion experiment. Given the RGB + depth images taken by 3 frames (discontinuous) RGB-D camera and the transformation matrix between them (taking the first frame as the reference frame), please generate point clouds from the above 3 frames of RGB-D images respectively and fuse the final point cloud output.
Code framework and data link: https://pan.baidu.com/s/1YPXvvR5bsXWnMk0juxqvgQ Extraction code: 7son
Reference answer:
#include "slamBase.hpp"
int main( int argc, char** argv )
{
CAMERA_INTRINSIC_PARAMETERS cameraParams{517.0,516.0,318.6,255.3,5000.0};
int frameNum = 3;
vector<Eigen::Isometry3d, Eigen::aligned_allocator<Eigen::Isometry3d>> poses;
PointCloud::Ptr fusedCloud(new PointCloud()); // Pay attention to memory alignment!!!
string path = "./data/";
string cameraPosePath = path + "cameraTrajectory.txt";
readCameraTrajectory(cameraPosePath, poses);
for (int idx = 0; idx < frameNum; idx++)
{
string rgbPath = path + "rgb/rgb" + to_string(idx) + ".png";
string depthPath = path + "depth/depth" + to_string(idx) + ".png";
FRAME frm;
frm.rgb = cv::imread(rgbPath);
if (frm.rgb.empty()) {
cerr << "Fail to load rgb image!" << endl;
}
frm.depth = cv::imread(depthPath, -1);
if (frm.depth.empty()) {
cerr << "Fail to load depth image!" << endl;
}
if (idx == 0) // If it is the first frame, turn the first frame into a point cloud
{
fusedCloud = image2PointCloud(frm.rgb, frm.depth, cameraParams);
}
else // If it is not the first frame, add the current frame to the point cloud, that is, point cloud fusion
{
fusedCloud = pointCloudFusion( fusedCloud, frm, poses[idx], cameraParams );
}
}
pcl::io::savePCDFile( "./fusedCloud.pcd", *fusedCloud ); // Save point cloud
return 0;
}
slamBase.cpp mainly realizes the definition of several important functions
#include "slamBase.hpp"
PointCloud::Ptr image2PointCloud(Mat rgb, Mat depth, CAMERA_INTRINSIC_PARAMETERS camera)
{
PointCloud::Ptr cloud ( new PointCloud );
for (int m = 0; m < depth.rows; m++)
for (int n = 0; n < depth.cols; n++)
{
// Gets the value at (m,n) in the depth map
ushort d = depth.ptr<ushort>(m)[n];
// d may not have a value, if so, skip this point
if (d == 0)
continue;
// If there is a value of d, add a point to the point cloud
PointT p;
// Calculate the spatial coordinates of this point
p.z = double(d) / camera.scale;
p.x = (n - camera.cx) * p.z / camera.fx;
p.y = (m - camera.cy) * p.z / camera.fy;
// Get its color from rgb image
p.b = rgb.ptr<uchar>(m)[n * 3];
p.g = rgb.ptr<uchar>(m)[n * 3 + 1];
p.r = rgb.ptr<uchar>(m)[n * 3 + 2];
// Add p to the point cloud
cloud->points.push_back(p);
}
// Set and save point clouds
cloud->height = 1;
cloud->width = cloud->points.size();
cloud->is_dense = false;
return cloud;
}
PointCloud::Ptr pointCloudFusion( PointCloud::Ptr &original, FRAME& newFrame, Eigen::Isometry3d T, CAMERA_INTRINSIC_PARAMETERS camera )
{
// ----------Start your code -------------//
// Simple point cloud overlay and fusion
PointCloud::Ptr newCloud(new PointCloud()), transCloud(new PointCloud());
newCloud = image2PointCloud(newFrame.rgb, newFrame.depth,camera); // At this time, the point cloud has pixel information, and the position xyz is the coordinate under the camera coordinate system
pcl::transformPointCloud(*newCloud,*transCloud,T.matrix()); // Convert the new point cloud from the camera coordinate system to the world coordinate system
*original += *transCloud; // Update point cloud
return original;
// ----------End your code -------------//
}
void readCameraTrajectory(string camTransFile, vector<Eigen::Isometry3d, Eigen::aligned_allocator<Eigen::Isometry3d>> &poses) // Pay attention to memory alignment
{
ifstream fcamTrans(camTransFile);
if(!fcamTrans.is_open())
{
cerr << "trajectory is empty!" << endl;
return;
}
// ----------Start your code -------------//
// Refer to job 8 to draw the track
string lineData;
while((getline(fcamTrans,lineData)) && !lineData.empty()){
Eigen::Quaterniond quad;
Eigen::Vector3d t;
Eigen::Isometry3d T = Eigen::Isometry3d::Identity();
if(lineData[0] == '#'){
continue;
}
istringstream strData(lineData);
strData>>t[0]>>t[1]>>t[2]>>quad.x()>>quad.y()>>quad.z()>>quad.w();
T.rotate(quad);
T.pretranslate(t);
//cout<<"test "<<endl;
poses.push_back(T);
}
// ----------End your code -------------//
}
Of course, there are header files:
# pragma once / / ensure that the header file is compiled only once
#include <iostream>
// opencv
#include <opencv2/core/core.hpp>
#include <opencv2/highgui/highgui.hpp>
// pcl
#include <pcl/point_types.h>
#include <pcl/io/pcd_io.h>
#include <pcl/common/transforms.h>
using namespace std;
using namespace cv;
typedef pcl::PointXYZRGB PointT;
typedef pcl::PointCloud<PointT> PointCloud;
// camera instrinsic parameters
struct CAMERA_INTRINSIC_PARAMETERS
{
double fx, fy, cx, cy, scale;
};
struct FRAME
{
cv::Mat rgb, depth;
};
PointCloud::Ptr image2PointCloud(Mat rgb, Mat depth, CAMERA_INTRINSIC_PARAMETERS camera);
PointCloud::Ptr pointCloudFusion( PointCloud::Ptr &original, FRAME& newFrame, Eigen::Isometry3d T, CAMERA_INTRINSIC_PARAMETERS camera );
void readCameraTrajectory(string camTransFile, vector<Eigen::Isometry3d, Eigen::aligned_allocator<Eigen::Isometry3d>> &poses);
Project structure
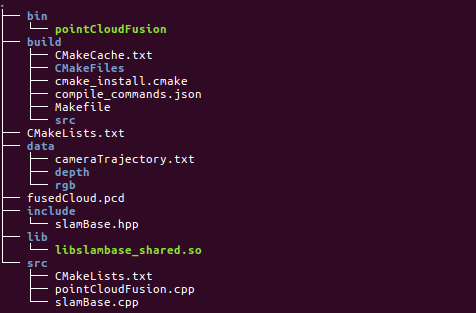
The project structure of this exercise is worth learning from. Put different files in different folders and be organized clearly.
Learn about cmakelists Txt:
First cmakelists txt:
CMAKE_MINIMUM_REQUIRED( VERSION 2.8 ) #Set version
PROJECT(PointCloud) #Set project name
SET( CMAKE_CXX_FLAGS "-std=c++11" )#Set compiler
SET(CMAKE_BUILD_TYPE Debug)
#Set the directory of executable binaries
SET( EXECUTABLE_OUTPUT_PATH ${PROJECT_SOURCE_DIR}/bin)
#Set the directory where the compiled library files are stored
SET( LIBRARY_OUTPUT_PATH ${PROJECT_SOURCE_DIR}/lib)
#Set header file directory
INCLUDE_DIRECTORIES( ${PROJECT_SOURCE_DIR}/include)
#And set the directory as the connection directory
LINK_DIRECTORIES( ${PROJECT_SOURCE_DIR}/lib)
#Add a subfolder, that is, enter the source code folder to continue the construction
ADD_SUBDIRECTORY( ${PROJECT_SOURCE_DIR}/src)
The second cmakelists Txt, continue to the src folder for construction:
# Increase the dependence of PCL Library
FIND_PACKAGE( PCL 1.7 REQUIRED )
FIND_PACKAGE( OpenCV REQUIRED )
ADD_DEFINITIONS( ${PCL_DEFINITIONS} )
INCLUDE_DIRECTORIES( ${PCL_INCLUDE_DIRS} )
LINK_LIBRARIES( ${PCL_LIBRARY_DIRS} )
INCLUDE_DIRECTORIES( ${PROJECT_SOURSE_DIR}/include )
ADD_LIBRARY(slambase_shared SHARED slamBase.cpp)
TARGET_LINK_LIBRARIES(slambase_shared
${OpenCV_LIBS}
${PCL_LIBRARIES} )
ADD_EXECUTABLE( pointCloudFusion pointCloudFusion.cpp )
TARGET_LINK_LIBRARIES(pointCloudFusion
slambase_shared
${OpenCV_LIBS}
${PCL_LIBRARIES}
)
Discussion:
How does the camera pose data stored in the camera trajectory file come from?
In monocular SLAM, feature points are usually extracted first, then the two images are matched and filtered, and finally the matched feature points are used to calculate the basic moment
Matrix or eigenmatrix, and then the camera pose can be estimated. At the same time, the three-dimensional points corresponding to the feature points can be triangulated, which is called map points. Then other frames are continuously added. The pose is calculated by PNP method, the three-dimensional points are maintained, and the pose is optimized by BA.
- Eigen memory alignment problem
When running the point cloud fusion experiment, an error was encountered during compilation:
/usr/include/eigen3/Eigen/src/Core/DenseStorage.h:128: Eigen::internal::plain_array<T, Size, MatrixOrArrayOptions, 32>::plain_array() [with T = double; int Size = 16; int MatrixOrArrayOptions = 0]: Assertion (reinterpret_cast<size_t>(eigen_unaligned_array_assert_workaround_gcc47(array)) & (31)) == 0 && "this assertion is explained here: " "http://eigen.tuxfamily.org/dox-devel/group__TopicU... " **** READ THIS WEB PAGE !!! ****"' failed.
Then I followed the prompt and went to the Eigen document to check the cause of the error, saying that it was related to fixed size vectorizable Eigen objects. However, after debugging, I found that the error appeared in the readCameraTrajectory() function and in positions push_ This error will be reported when back (T) is stored in the third pose. And even:: isometriy3d checked, it should not belong to fixed size vectorizable even objects. Miraculously, when I delete the pose data of the third frame and only read the pose data of the first two frames, the code can be executed and generated smoothly pcd file.
This problem bothered me for two days,
terms of settlement:
Add the memory alignment parameter when initializing poses:
vector<Eigen::Isometry3d, Eigen::aligned_allocator<Eigen::Isometry3d>> poses And in readCameraTrajectory()(The reference and definition of the corresponding function calling the parameter are also poses Make the same changes
- pcl_viewer display black screen problem
This is a problem left over by history after I just installed pcl at that time. At that time, I installed pcl library according to several tutorials, which was called a tragic one. I don't know how many times I have reinstalled, and finally I can compile smoothly, but the only drawback is pcl_ There is something wrong with the viewer.
It may be that both pcl and pcl compiled by Ubuntu are installed at the same time, and pcl is being used_ When the viewer is, the point cloud cannot be rendered and the window is dark.
Loading fusedCloud.pcd [PCLVisualizer::setUseVbos] Has no effect when OpenGL version is ≥ 2 [done, 4361.73 ms : 706468 points]」
I always thought it was the version of opengl. Later, I found that there was a pcl under / usr/bin / and usr/local/bin respectively_ Viewer, the default execution is the one under / usr/local/bin. Try the one under / usr/bin / and suddenly it will display normally. (it seems that the built-in pcl in Ubuntu works).
These two epic pits took me three days, but they also forced me to turn over the Eigen library and increase my experience in dealing with mistakes.
Point Cloud Filtering
subject
Title: given a fused point cloud, please down sample it first, then filter it, and finally output the filtered results and filtered outliers.
Point cloud filtering code framework and pending data download: link: https://pan.baidu.com/s/1EAP89pHV-Ec8glKxaATBAQ Extraction code: as3x
#include <pcl/point_types.h>
#include <pcl/io/io.h>
#include <pcl/io/pcd_io.h>
#include <pcl/visualization/cloud_viewer.h>
#include <pcl/filters/radius_outlier_removal.h>
#include <pcl/filters/voxel_grid.h>
#include <pcl/filters/statistical_outlier_removal.h>
#include <boost/thread.hpp>
typedef pcl::PointXYZRGB PointT;
int main(int argc, char** argv)
{
// Load point cloud initialization
pcl::PointCloud<PointT>::Ptr cloud(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_downSampled(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_filtered(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr noise(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_filtered_R(new pcl::PointCloud<PointT>);
if (pcl::io::loadPCDFile("./data/fusedCloud.pcd", *cloud) == -1)
{
cout << "Point cloud data reading failed!" << endl;
}
std::cout << "Points number after filtering: " << cloud->points.size() << std::endl;
// -----------Start your code, refer to http://docs.pointclouds.org/trunk/group__filters.html --------
// Down sampling while maintaining the shape characteristics of point cloud
pcl::VoxelGrid<PointT> downSampled; // Create filter object
downSampled.setInputCloud(cloud); //Set the point cloud to be filtered to the filtered object
downSampled.setLeafSize(0.01f, 0.01f, 0.01f); // Set the voxel created during filtering to a cube with a side length of 1cm
downSampled.filter(*cloud_downSampled); // Perform filtering operation and store the output
// Statistical filtering
pcl::StatisticalOutlierRemoval<PointT> statisOutlierRemoval; // Filter object creation
statisOutlierRemoval.setInputCloud(cloud_downSampled); // Set filtered point cloud
statisOutlierRemoval.setMeanK(50); // Set the number of adjacent query points to be considered in statistics
statisOutlierRemoval.setStddevMulThresh(1.0); // Set the threshold to judge whether it is an outlier: mean + 1.0 × standard deviation
statisOutlierRemoval.filter(*cloud_filtered); // The filtering results are stored in cloud_filtered
// Try radius filtering
pcl::RadiusOutlierRemoval<PointT> RoutlierRemoval; //Create filter object
RoutlierRemoval.setInputCloud(cloud_downSampled); // Set the point cloud to be filtered
RoutlierRemoval.setRadiusSearch(0.02); // Set search radius
RoutlierRemoval.setMinNeighborsInRadius(2); // Set the minimum number of neighbors for an interior point
RoutlierRemoval.filter(*cloud_filtered_R);
std::cout << "Points number after filtering: " << cloud_filtered->points.size() << std::endl;
// Output interior point
pcl::PCDWriter writer;
writer.write<PointT>("./data/cloud_inliers.pcd", *cloud_filtered, false);
// Output outliers
statisOutlierRemoval.setNegative(true);
statisOutlierRemoval.filter(*noise);
writer.write<PointT>("./data/cloud_outliers.pcd", *noise, false);
// -----------End your code--------
// Display the filtering results, or use pcl_viewer command view
boost::shared_ptr<pcl::visualization::PCLVisualizer> viewer(new pcl::visualization::PCLVisualizer("3D Viewer"));
viewer->setBackgroundColor(0, 0, 0);
pcl::visualization::PointCloudColorHandlerRGBField<PointT> rgb(cloud_filtered);
viewer->addPointCloud<PointT> (cloud_filtered, rgb, "sample cloud");
while (!viewer->wasStopped())
{
viewer->spinOnce(100);
// boost::this_thread::sleep(boost::posix_time::microseconds(100000));
}
return (0);
}
Attached cmakelists Txt, which is similar to point cloud fusion. Here, only cmakelists built in src:
FIND_PACKAGE( PCL 1.7 REQUIRED )
FIND_PACKAGE( OpenCV REQUIRED )
ADD_DEFINITIONS( ${PCL_DEFINITIONS} )
INCLUDE_DIRECTORIES( ${PCL_INCLUDE_DIRS} )
LINK_LIBRARIES( ${PCL_LIBRARY_DIRS} )
ADD_EXECUTABLE( pointCloudFilter pointCloudFilter.cpp )
TARGET_LINK_LIBRARIES(pointCloudFilter
${OpenCV_LIBS}
${PCL_LIBRARIES}
${Boost_LIBRARIES}
)
discuss
- Voxel
Voxel is the abbreviation of Volume Pixel, and the solid containing voxel can pass through volume rendering Or extract a given threshold Outline polygon Isosurface Show it. As its name suggests, digital data three-dimensional space The smallest unit in segmentation. Voxels are used for three-dimensional imaging, scientific data and Medical imaging And other fields. Conceptually, it is similar to the smallest unit of two-dimensional space - pixels, which are used in the image data of two-dimensional computer images. Some real 3D displays use voxels to describe their resolution. For example, they can display 512 × five hundred and twelve × 512 voxel display.
- Down sampling
The main purposes of reducing the image (or called subsampled or downsampled) are two: 1. Making the image conform to the size of the display area; 2. Generate thumbnails of corresponding images. The main purpose of enlarging an image (or called upsampling or image interpolation) is to enlarge the original image so that it can be displayed on a higher resolution display device. The scaling operation of the image can not bring more information about the image, so the quality of the image will inevitably be affected. However, there are some scaling methods that can increase the information of the image, so that the quality of the scaled image exceeds that of the original image.
It feels like the convolution in the process of CNN.
pcl provides the down sampling class, template < typename pointt > class pcl:: voxelgrid < pointt >
The principle is to use the centroid of a voxel as the approximate value of all points in the voxel range.
- Statistical filtering
Make a statistical analysis of the neighborhood of each point, and trim some points that do not meet the standard. The specific method is to calculate the distance distribution from the point to the adjacent point in the input data. For each point, calculate the average distance from it to all adjacent points (assuming that the result is a Gaussian distribution, and its shape is determined by the mean and standard deviation). Then the points with the average distance outside the standard range can be defined as outliers and removed from the data.
- Radius filtering
As shown in the figure below, it is helpful to visually understand the role of RadiusOutlierRemoval. In point cloud data, the user specifies that there should be at least enough close neighbors around each point within a certain range. For example, if you specify at least 1 neighbor, only yellow points will be deleted. If you specify at least 2 neighbors, both yellow and green points will be deleted.
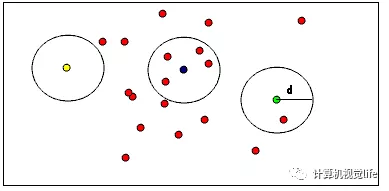
In addition, another bug was encountered. During compilation, there was an error in boost:
CMakeFiles/pointCloudFilter.dir/pointCloudFilter.cpp.o: In function'boost::this_thread::sleep(boost::posix_time::ptime const&)'Medium: /usr/include/boost/thread/pthread/thread_data.hpp:278: yes'boost::this_thread::hiden::sleep_until(timespec const&)'Undefined reference collect2: error: ld returned 1 exit status
Not solved... But the function of this sentence is sleep delay. It's harmless to remove it (personally).
Point cloud smoothing
subject
Title: given a fused point cloud, it has been down sampled and filtered (the code has been given). Smooth it (output the result) and calculate
Normal, and the normal is displayed on the smoothed point cloud.
Code frame link: https://pan.baidu.com/s/1zbCA8WgE0PUD4eqDyxFrkw Extraction code: j7zc
reference material: Learn SLAM | point cloud smoothing normal estimation from scratch
Knowledge points: smoothing and normal estimation
Reference results: if everything goes well, you will get the attached results. You can adjust the density of the normal and zoom in to see whether the normal calculation meets the expectation

The code is based on the previous job, that is, smoothing the filtered point cloud, mainly using resampling.
#include <pcl/point_types.h>
#include <pcl/io/io.h>
#include <pcl/io/pcd_io.h>
#include <pcl/visualization/cloud_viewer.h>
#include <pcl/filters/radius_outlier_removal.h>
#include <pcl/filters/voxel_grid.h>
#include <pcl/filters/statistical_outlier_removal.h>
#include <pcl/surface/mls.h>
#include <pcl/features/normal_3d.h>
typedef pcl::PointXYZRGB PointT;
int main(int argc, char** argv)
{
// Load input file
pcl::PointCloud<PointT>::Ptr cloud(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_downSampled(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_filtered(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_smoothed(new pcl::PointCloud<PointT>);
if (pcl::io::loadPCDFile("./data/fusedCloud.pcd", *cloud) == -1)
{
cout << "Point cloud data reading failed!" << endl;
}
std::cout << "Orginal points number: " << cloud->points.size() << std::endl;
// Down sampling while maintaining the shape characteristics of point cloud
pcl::VoxelGrid<PointT> downSampled; //Filter object creation
downSampled.setInputCloud (cloud); //Set the point cloud to be filtered to the filtered object
downSampled.setLeafSize (0.01f, 0.01f, 0.01f); //A cube with a voxel volume of 1cm created during filtering is set
downSampled.filter (*cloud_downSampled); //Perform filtering processing and store the output
pcl::io::savePCDFile ("./downsampledPC.pcd", *cloud_downSampled);
// Statistical filtering
pcl::StatisticalOutlierRemoval<PointT> statisOutlierRemoval; //Create filter object
statisOutlierRemoval.setInputCloud (cloud_downSampled); //Set the point cloud to be filtered
statisOutlierRemoval.setMeanK (50); //Set the number of adjacent query points to be considered in statistics
statisOutlierRemoval.setStddevMulThresh (3.0); //Set the threshold to judge whether it is an outlier: mean + 1.0 * standard deviation
statisOutlierRemoval.filter (*cloud_filtered); //The filtering results are stored in cloud_filtered
pcl::io::savePCDFile ("./filteredPC.pcd", *cloud_filtered);
// ----------------------Start your code--------------------------//
// Please refer to PCL official website to realize the following functions
// Resampling point clouds
pcl::search::KdTree<PointT>::Ptr treeSampling(new pcl::search::KdTree<PointT>); // Create nearest neighbor KD tree
pcl::MovingLeastSquares<PointT,PointT> mls_filter; // Defines the object of the least squares implementation
mls_filter.setInputCloud(cloud_filtered); // Input point cloud to be processed
mls_filter.setComputeNormals(false); // Sets whether calculated normals need to be stored in the least squares calculation
mls_filter.setPolynomialOrder(2); // Second order polynomial fitting
mls_filter.setSearchMethod(treeSampling); //Set KD tree as the search method
mls_filter.setSearchRadius(0.05); //Sets the K-nearest neighbor radius for fitting
mls_filter.process(*cloud_smoothed); //output
// Output resampling results
pcl::io::savePCDFile("./smoothedPC.pcd", *cloud_smoothed);
// Normal estimation
pcl::NormalEstimation<PointT, pcl::Normal> normalEstimation; // Create objects with normal estimates
normalEstimation.setInputCloud(cloud_smoothed); // Input point cloud
pcl::search::KdTree<PointT>::Ptr tree(new pcl::search::KdTree<PointT>); //Create KD tree
normalEstimation.setSearchMethod(tree);
pcl::PointCloud<pcl::Normal>::Ptr normals(new pcl::PointCloud<pcl::Normal>); //Define cloud normals
normalEstimation.setKSearch(10); // Use the nearest 10 points around the current point
//normalEstimation.setRadiusSearch(0.03);
normalEstimation.compute(*normals); // Calculate normals
// ----------------------End your code--------------------------//
// Display results
boost::shared_ptr<pcl::visualization::PCLVisualizer> viewer (new pcl::visualization::PCLVisualizer ("PCL Viewer"));
viewer->setBackgroundColor (0, 0, 0);
pcl::visualization::PointCloudColorHandlerRGBField<PointT> rgb(cloud_smoothed);
viewer->addPointCloud<PointT> (cloud_smoothed, rgb, "smooth cloud");
viewer->setPointCloudRenderingProperties (pcl::visualization::PCL_VISUALIZER_POINT_SIZE, 3, "smooth cloud");
viewer->addPointCloudNormals<PointT, pcl::Normal> (cloud_smoothed, normals, 20, 0.05, "normals");
viewer->initCameraParameters ();
while (!viewer->wasStopped ())
{
viewer->spinOnce (100);
// boost::this_thread::sleep (boost::posix_time::microseconds (100000));
}
return 1;
}
CMakeLists.txt, like the previous exercises, is nothing more than the introduction of PCL and OpenCV libraries.
discuss
Mainly resampling and normal estimation:
Point cloud resampling is actually realized through a method called "Moving Least Squares" (MLS), and the corresponding class name is: PCL:: movingleast squares.
- MLS
The moving least square method is improved on the basis of the least square method, and the weight function is added. For details, please refer to the paper, "Moving least squares paper" link , this paper explains MLS in detail, and finally gives the idea of program design.
- Normal estimation
Normals are very useful, especially widely used in 3D modeling. For example, in the field of computer graphics, normals determine the Flat Shading of surfaces and light sources. For each point light source position, its brightness depends on the direction of surface normals.
Here, the surface normal is directly inferred from the point cloud data set by using the approximate value, and the surface normal of each point in the point cloud is approximately estimated.
Specifically, the problem of estimating the surface normal of a point is simplified to the analysis of the eigenvector and eigenvalue of the covariance matrix calculated from the nearest neighbor of the point, which will not be introduced here. PCL has encapsulated the function for us.
K-Neighborhood: it is necessary to estimate the surface normal at a point from the neighborhood around the point (also known as K-Neighborhood), so the selection of this K-Neighborhood is also crucial
- K-D-tree
KD tree is a data structure and a special case of spatial binary tree. It can be easily used for range search. KD tree is used here to facilitate the management and search of point cloud. This structure can easily find the nearest neighbor.
K-d tree (k-dimensional tree for short) is a data structure that divides k-dimensional data space. It is mainly applied to the search of key data in multidimensional space (such as range search and nearest neighbor search). K-d tree is a special case of binary space partition tree.)
Here is a 3-tree:
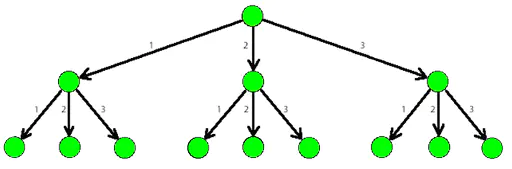
k-d has advantages in neighborhood search, but in the case of large amount of data, if the partition granularity is small, the cost of building a tree is also large, but it is more flexible than octree. In general, the search efficiency will increase when the amount of data is small, but it will decrease when the search efficiency is high.
reference resources: https://www.cnblogs.com/zfyouxi/p/4795584.html
Point cloud gridding
subject
Title: given the input point cloud, filter and smooth the point cloud in combination with the previous content, and calculate the normal. Finally, mesh it with greedy projection triangulation method to display the meshing results.
Code frame: link: https://pan.baidu.com/s/1avSGdi4IG3ry3wNCI_jDLQ Extraction code: cxjy
Knowledge points: greedy projection triangulation method
#include <pcl/point_types.h>
#include <pcl/io/io.h>
#include <pcl/io/pcd_io.h>
#include <pcl/visualization/cloud_viewer.h>
#include <pcl/filters/radius_outlier_removal.h>
#include <pcl/filters/voxel_grid.h>
#include <pcl/filters/statistical_outlier_removal.h>
#include <pcl/surface/mls.h>
#include <pcl/features/normal_3d.h>
#include <pcl/surface/gp3.h>
#include <pcl/surface/poisson.h>
typedef pcl::PointXYZ PointT;
int main(int argc, char** argv)
{
// Load input file
pcl::PointCloud<PointT>::Ptr cloud(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_downSampled(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_filtered(new pcl::PointCloud<PointT>);
pcl::PointCloud<PointT>::Ptr cloud_smoothed(new pcl::PointCloud<PointT>);
if (pcl::io::loadPCDFile("./data/fusedCloud.pcd", *cloud) == -1)
{
cout << "Point cloud data reading failed!" << endl;
}
std::cout << "Orginal points number: " << cloud->points.size() << std::endl;
// ----------------------Start your code--------------------------//
// Please refer to the previous article on point cloud down sampling, filtering, smoothing and other contents, as well as the PCL official website to achieve the following functions. The code is not difficult.
// Down sampling noise reduction
pcl::VoxelGrid<PointT> downSampled; //Filter object creation
downSampled.setInputCloud(cloud);
downSampled.setLeafSize(0.01f,0.01f,0.01f);
downSampled.filter(*cloud_downSampled);
pcl::io::savePCDFile("./downsampled.pcd", *cloud_downSampled);
// Statistical filtering noise reduction again
pcl::StatisticalOutlierRemoval<PointT> statisOutlierRemoval;
statisOutlierRemoval.setInputCloud(cloud_downSampled);
statisOutlierRemoval.setMeanK(50); // Set the number of adjacent query points to be considered in statistics
statisOutlierRemoval.setStddevMulThresh(1.0); // Set the threshold to judge whether it is an outlier: mean + 1.0 × standard deviation
statisOutlierRemoval.filter(*cloud_filtered);
pcl::io::savePCDFile("./filteredPC.pcd", *cloud_filtered);
// Resampling point clouds
pcl::search::KdTree<PointT>::Ptr treeSampling(new pcl::search::KdTree<PointT>);
pcl::PointCloud<PointT> mls_points; //Output MLS
pcl::MovingLeastSquares<PointT, PointT> mls; // Define least squares implementation object mls
mls.setInputCloud(cloud_filtered);
mls.setComputeNormals(false); // Sets whether calculated normals need to be stored in the least squares calculation
mls.setPolynomialOrder(2); // Second order polynomial fitting
// mls.setPolynomialFit(false); // Set to false to speed up smooth. The new version is not recommended
mls.setSearchMethod(treeSampling); // Set search method
mls.setSearchRadius(0.05); // Unit: m sets the K-nearest neighbor radius used for fitting
mls.process(mls_points); // The output can also be directly output to cloud_smoothed, the following transformation is to experiment with these two data structures
// mls results are first transformed into pointcloud2, and then back to the experiment
pcl::PCLPointCloud2 tmp;
pcl::toPCLPointCloud2(mls_points, tmp);
pcl::fromPCLPointCloud2(tmp, *cloud_smoothed);
pcl::io::savePCDFile("./resampledPC.pcd", mls_points);
std::cout<<"Points number after filtering and resampling:"<<cloud_smoothed->points.size()<<std::endl;
// Normal estimation
pcl::NormalEstimation<PointT, pcl::Normal> normalEstimation;
normalEstimation.setInputCloud(cloud_smoothed);
pcl::search::KdTree<PointT>::Ptr tree(new pcl::search::KdTree<PointT>);
normalEstimation.setSearchMethod(tree);
pcl::PointCloud<pcl::Normal>::Ptr normals(new pcl::PointCloud<pcl::Normal>); // Defines the output normal of the point cloud
// The K-nearest neighbor determination method uses K nearest points or determines the point set of a circle with r as the radius. Both methods can take 1
normalEstimation.setKSearch(10); // Use the nearest 10 points around the current point
// normalEstimation.setRadiusSearch(0.03) // For each point, the nearest neighbor search method with a radius of 3cm is used
normalEstimation.compute(*normals); // Calculate normals
// Connect the pose and normal information of point cloud together, that is, PointNormal type, directed point cloud
pcl::PointCloud<pcl::PointNormal>::Ptr cloud_with_normals(new pcl::PointCloud<pcl::PointNormal>);
pcl::concatenateFields(*cloud_smoothed, *normals,*cloud_with_normals);
pcl::PolygonMesh mesh; // Storage final triangulated network model
// Define search tree objects
pcl::search::KdTree<pcl::PointNormal>::Ptr tree2(new pcl::search::KdTree<pcl::PointNormal>);
tree2->setInputCloud(cloud_with_normals);
// Greedy projection triangulation
pcl::GreedyProjectionTriangulation<pcl::PointNormal> gp3; // Define triangulated objects
// Set triangulation parameters
gp3.setSearchRadius(0.1); // Set the radius when searching, that is, the radius of KNN
gp3.setMu(2.5); // Set the maximum distance of sample points to search their adjacent points to 2.5 times (typical value is 2.5-3), which makes the algorithm use the change of point cloud density
gp3.setMaximumNearestNeighbors(100); // Set the maximum number of neighborhoods that can be searched for sample points, typically 50-100
gp3.setMinimumAngle(M_PI/18); // Set the minimum angle of triangle internal angle after triangulation to 10
gp3.setMaximumAngle(2*M_PI/3); //After setting the triangle lake, the inner angle of the triangle is 120
gp3.setMaximumSurfaceAngle(M_PI/4); // Set the maximum angle that the normal direction of a point deviates from the normal of the sample point to 45. If it exceeds, the point will not be considered during connection
gp3.setNormalConsistency(true); // Set this parameter to ensure that the normal orientation is consistent. If it is set to false, the normal consistency check will not be carried out
gp3.setInputCloud(cloud_with_normals); // Set the input point cloud as a directed point cloud
gp3.setSearchMethod(tree2); // Set search method
gp3.reconstruct(mesh); // Reconstruction extraction triangulation
// ----------------------End your code--------------------------//
// Show gridding results
boost::shared_ptr<pcl::visualization::PCLVisualizer> viewer(new pcl::visualization::PCLVisualizer("3D Viewer"));
viewer->setBackgroundColor(0, 0, 0); //
viewer->addPolygonMesh(mesh, "mesh"); //
while (!viewer->wasStopped())
{
viewer->spinOnce(100);
//boost::this_thread::sleep(boost::posix_time::microseconds(100000));
}
return 1;
}
discuss
There are still a lot of contents about grid, which can be written by brother Liu Make series together , time is limited, so we will not study its principle deeply. When we have time in the future, we can play this module well.
**Suggestion: * * greedy projection triangulation used is not commonly used. There are many Poisson reconstruction methods actually used. You can try them
See the difference.
**Q: * * what is a directed point cloud? Aren't all the point clouds obtained in practice disordered? How to mesh the ordered point cloud and the disordered point cloud?
A: ordered point cloud refers to that the point clouds are arranged in order, which can be understood as an index method similar to 2D images, so it is easy to find its adjacent point information. However, in general, the point clouds we get are unorganized. Whether ordered or disordered point clouds, there is no difference in meshing. Gridding needs to find neighborhood information. The ordered point cloud itself has information. For the disordered point cloud, we will create objects such as kdtree during gridding. The purpose is to find the neighborhood for indexing. In short, it can be considered that there is no difference in use.
For function setPolynomialFit(false)
mls.setPolynomialFit(false); // Set to false to speed up smooth* // Warning when using 'void pcl::MovingLeastSquares<PointInT, PointOutT>::setPolynomialFit(bool) [with PointInT = pcl::PointXYZ; PointOutT = pcl::PointXYZ]' is deprecated: use setPolynomialOrder() instead (It will be removed in PCL 1.12) [-Wdeprecated-declarations]
As you can see, this is not recommended. This function is used in pcl1 12 will be removed (PCL1.10 I currently use). According to the description, the function should be and setPolynomialOrder() instead of??? When using, you can comment out this code directly.
reference resources:
Learn SLAM from scratch | Hello, point cloud
Learn SLAM | point cloud smoothing normal estimation from scratch
Learn from scratch the evolution of SLAM | point cloud to grid